Archives
- 2026-05
- 2026-04
- 2026-03
- 2026-02
- 2026-01
- 2025-12
- 2025-11
- 2025-10
- 2025-09
- 2025-03
- 2025-02
- 2025-01
- 2024-12
- 2024-11
- 2024-10
- 2024-09
- 2024-08
- 2024-07
- 2024-06
- 2024-05
- 2024-04
- 2024-03
- 2024-02
- 2024-01
- 2023-12
- 2023-11
- 2023-10
- 2023-09
- 2023-08
- 2023-06
- 2023-05
- 2023-04
- 2023-03
- 2023-02
- 2023-01
- 2022-12
- 2022-11
- 2022-10
- 2022-09
- 2022-08
- 2022-07
- 2022-06
- 2022-05
- 2022-04
- 2022-03
- 2022-02
- 2022-01
- 2021-12
- 2021-11
- 2021-10
- 2021-09
- 2021-08
- 2021-07
- 2021-06
- 2021-05
- 2021-04
- 2021-03
- 2021-02
- 2021-01
- 2020-12
- 2020-11
- 2020-10
- 2020-09
- 2020-08
- 2020-07
- 2020-06
- 2020-05
- 2020-04
- 2020-03
- 2020-02
- 2020-01
- 2019-12
- 2019-11
- 2019-10
- 2019-09
- 2019-08
- 2019-07
- 2019-06
- 2019-05
- 2019-04
- 2018-07
-
Artificial permutations have also been exploited by protein
2020-12-26
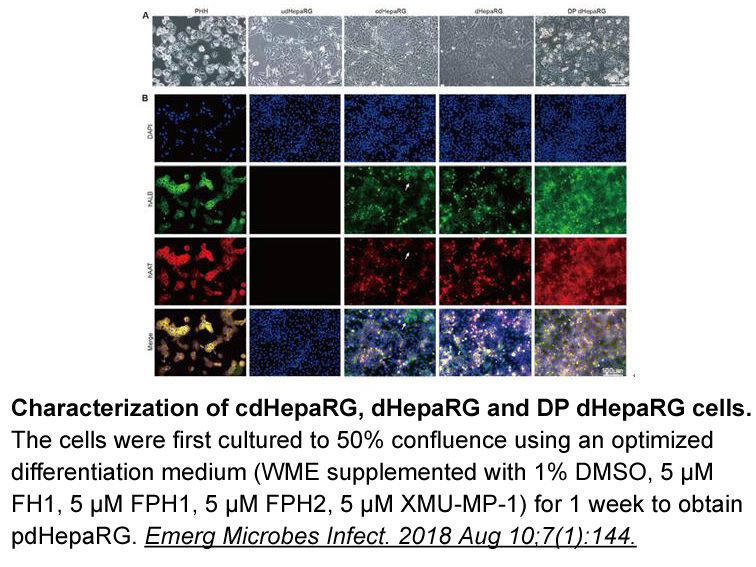
Artificial permutations have also been exploited by protein engineering to manipulate protein scaffolds, in order to improve catalytic activity, alter substrate or ligand binding affinity, reduce proteolytic susceptibility, increase stability, generate different aggregation states and improve fluore
-
In summary we investigated the fluoride sensitivity of
2020-12-26
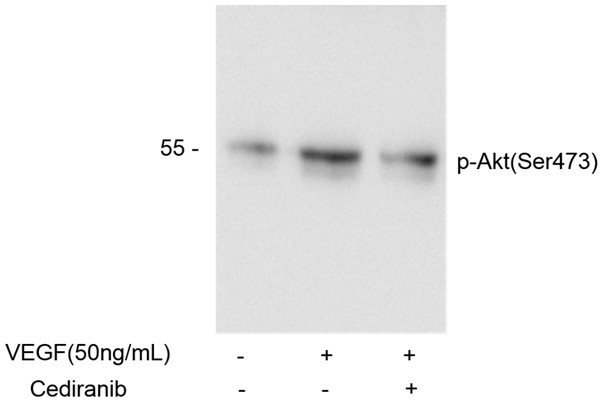
In summary, we investigated the fluoride sensitivity of different S. mutans strains in terms of enolase activity. Lower enolase activity was not always associated with lower S. mutans growth in cultures with NaF. Gene analysis showed that UA130 and NCH105 both have enolase point mutations. Unique am
-
All known vertebrate TRIMs are categorized
2020-12-25
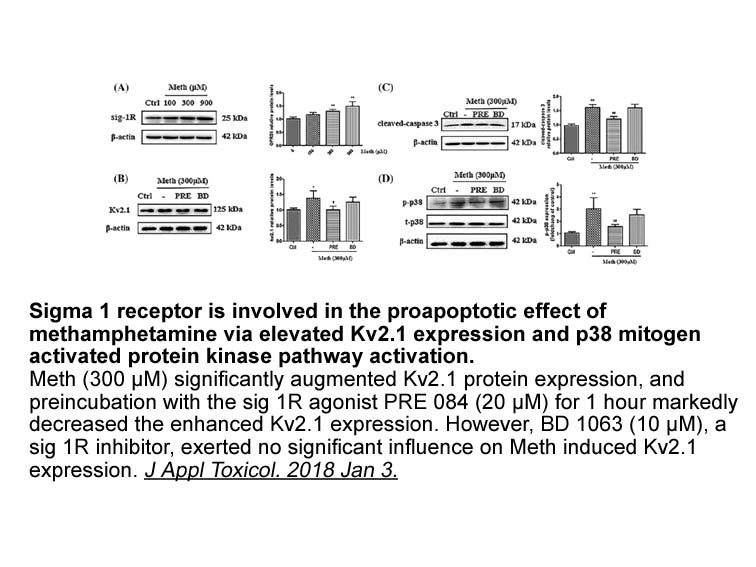
All known vertebrate TRIMs are categorized in 11 distinct subclasses depending on the types of domains present at their carboxyl-terminals (Fig. 3) [29], [35]. Beyond conserved N-terminal domains, it is the C-terminal that provides specificity of interactions with other proteins. The subclass IV for
-
br RING type E s and their substrates There
2020-12-25
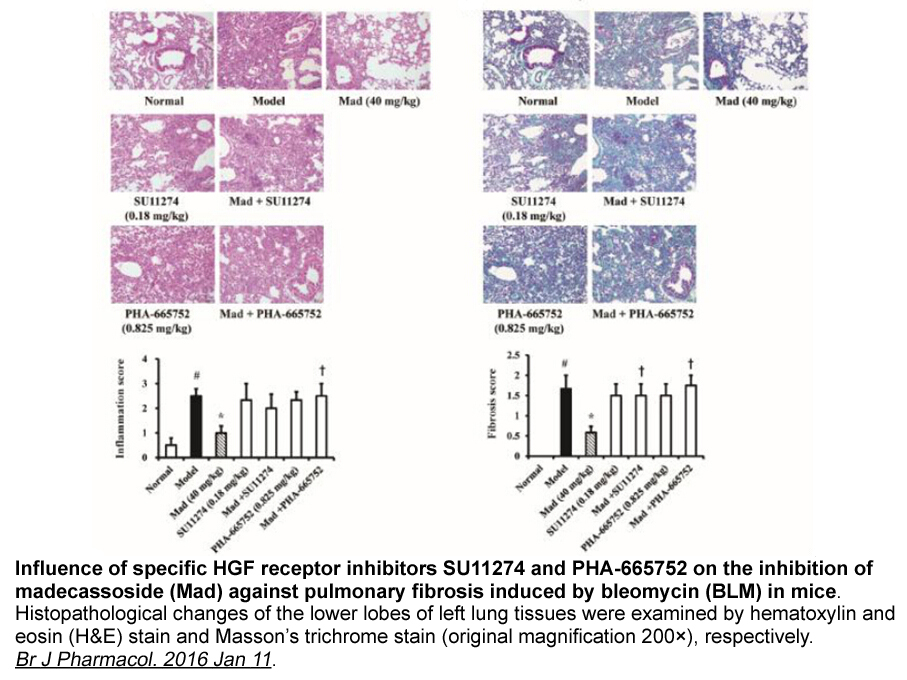
RING-type E3s and their substrates There is enormous diversity in substrate ubiquitination and its regulation, as the targets of RING-type E3s are incredibly varied. RING-type E3s are implicated as tumor suppressors, oncogenes, and mediators of endocytosis, and play critical roles in complex mult
-
Pyruvate dehydrogenase multienzyme complex PDHc catalyzes th
2020-12-25
Pyruvate dehydrogenase multienzyme complex (PDHc) catalyzes the oxidative decarboxylation of pyruvate, and subsequently acetylates coenzyme A (CoA) to acetyl-CoA during the tricarboxylic MAFP metabolic pathway using thiamine diphosphate (ThDP) and Mg2+ as cofactors. PDHc poses a key role in cyanoba
-
br DNA PK After sensing and binding to the DSB
2020-12-25
DNA-PK After sensing and binding to the DSB, Ku quickly recruits DNA-PKcs to the site of the DNA break. Similar to Ku70/80, recruitment of DNA-PKcs to DSBs occurs within seconds of their creation [12]. The interaction between Ku70/80 and DNA-PKcs requires the presence of dsDNA and the complex for
-
Additional derivatives were synthesised using
2020-12-25
Additional derivatives were synthesised using a modified approach (). Commercially available 4-bromo-2-methoxyaniline () was converted into the boronic ester , followed by a Suzuki–Miyaura cross-coupling with chromenone triflate to afford the corresponding arylamine . Acylation of with chloroacety
-
br Structural and biochemical features of pol X family polym
2020-12-25
Structural and biochemical features of pol X family polymerases The X-family DNA polymerase (polX) is specialized in DNA repair. This family is composed of four different DNA polymerases: pol β, pol λ, pol μ and TdT. Only three members of the polX family possess an N-terminal BRCT (BRCA1 carboxy-
-
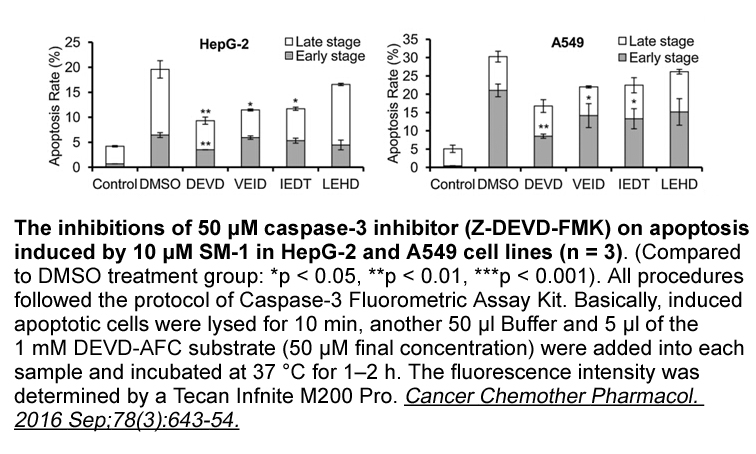
The inhibition of DNMT using
2020-12-25
The inhibition of DNMT1 using 5-AZA-dC or DNMT1 siRNA increased TIMP2 protein and mRNA expression, suggesting that TIMP2 expression is down-regulated by DNA methylation in the HDFs. In addition, 5-AZA-dC treatment led to a dose-dependent decrease of DNMT1 protein expression. 5-AZA-dC is a cytosine a
-
Except for S adenosylmethionine SAM
2020-12-25
Except for S-adenosylmethionine (SAM, Fig. 1), sources of endogenous DNA alkylation are not well defined. Other possible sources include nitroso compounds related to the well known mutagen methylnitrosourea which are generated in vitro by nitrosation of cellular amines including amino acids, protein
-
The most significant source of DAG originates from the
2020-12-25
The most significant source of DAG originates from the PLC family of enzymes, which produces DAG in an agonist-regulated reaction. PLCs constitute a wide family of enzymes acting on both phosphatidilinositols (PI) and phosphatidilcoline (PC) (Li et al., 2010) in the plasma membrane and other intrace
-
br Mammalian DGKs current knowledge Despite our understandin
2020-12-25

Mammalian DGKs: current knowledge Despite our understanding of prokaryotic DGKs, our limited understanding of the structure of mammalian DGKs is a major gap in our knowledge. Importantly, major differences exist between prokaryotic and mammalian enzymes often revealing the fact that structural fe
-
br Conclusions This report describes the discovery of a
2020-12-25
Conclusions This report describes the discovery of a new class of host-directed antiviral agents characterized by a 1-aryl-4,6-diamino-1,2-dihydrotriazine scaffold, responsible for a host (human) DHFR inhibition mechanism. Host-targeting antivirals represent an alternative and emerging strategy t
-
As shown in Fig there are
2020-12-25
As shown in Fig. 9, there are two mechanisms for the removal of the Va-acyl group from PC to make it available for incorporation into TAG with DGATs' acting at the final acylation step. ①: Transfer of Va from PC to the acyl-CoA pool. This process can be driven by the reverse action of acyl- CoA:lyso
-
Reports on the participation of NDH along with other
2020-12-25
Reports on the participation of NDH2 along with other respiratory complexes in the formation of a large supercomplex (Grandier-Vazeille et al. 2001) or even a respirosome-like structure in S. cerevisiae mitochondria (Matus-Ortega et al. 2015) are very intriguing. Similarly, in Y. lypolytica mitochon
15930 records 740/1062 page Previous Next First page 上5页 736737738739740 下5页 Last page